BDB-Lab March 2026 Updates
What lurks in urban soil microbiomes
Looking Forward. Luis will present “Big data and small genes. The small proteins of the global microbiome” at the 55th Annual Meeting of SBBq in Águas de Lindóia, São Paulo, Brazil (May 16-19).
Focus of the Quarter: long-read metagenomics on urban soil by Yiqian Duan
This urban soil project, initially supported by EMBARK and now by SEARCHER, was initiated to explore the distribution of antibiotic resistance genes (ARGs) in urban soil environments using long-read sequencing. Between May and August 20231, we collected a total of 58 urban soil samples from university campuses and parks in two major cities (Shanghai and Nanjing) of China. Each sample was subjected to deep sequencing, resulting in a minimum of 40 Gbp of both short-read and long-read data per sample.
During the subsequent analysis, we found that the long-read sequencing data from urban soils provided insights beyond the ARGs that were our original focus. For instance, it enabled the recovery of more complete and less contaminated microbial genomes, which facilitated the discovery of novel microbial species as well as new functional genes and gene clusters. This has significantly expanded our understanding of urban soil microbial communities. The main results have now been released as a preprint:
We obtained 7,949 medium- and high-quality metagenome-assembled genomes (MAGs), 1,060 of which were near-finished quality (high-quality regarding MIMAG criteria) and highly contiguous. Collectively, these MAGs comprise 4,171 species-level genome bins, of which over 97% represent previously undescribed species.
We identified more than 30,000 biosynthetic gene clusters from MAGs assembled from long reads, which were highly contiguous compared with those derived from short-read assemblies.
We uncovered over 2 million small protein families, associated with defense systems and mobile genetic elements, highlighting their overlooked roles in the urban soil.
We found a large repertoire of latent antibiotic resistance genes (ARGs) and only a small number of established ARGs in urban soils.
All generated data and resources are freely available. If you want to explore the MAGs and access the associated data, you could visit our Urban Soil MAG website as well as the Zenodo repository.
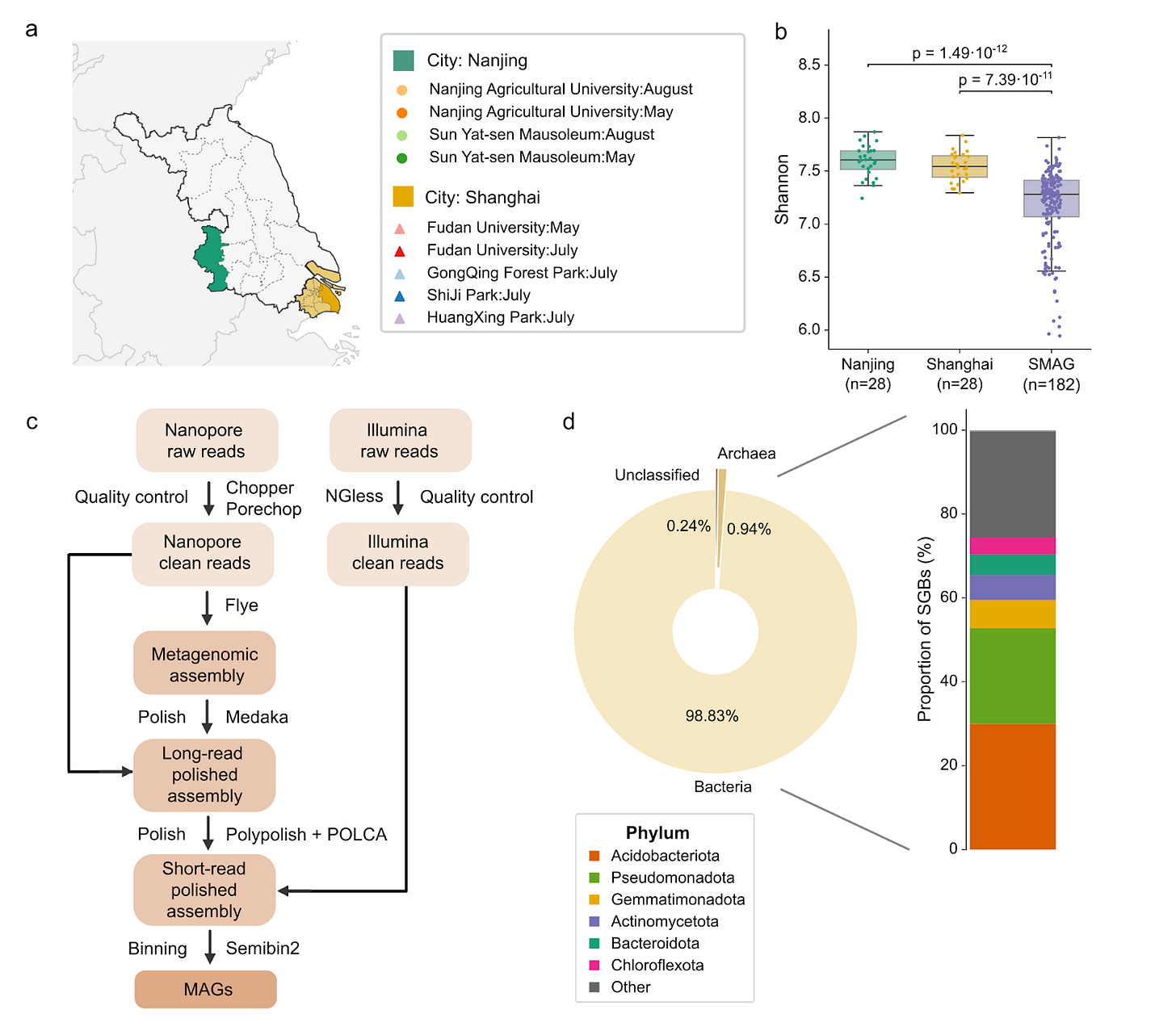
Urban soil microbiome using long-read metagenomics. (a) Graphical representation of samples from two major cities in eastern China, Nanjing and Shanghai, where samples were collected. (b) Comparison of the Shannon index between the two cities and samples from a publicly available soil catalog. (c) Workflow of assembly and binning using long and short reads. (d) Taxonomic classification rates of SGBs at different taxonomic levels.
Full citation:
Yiqian Duan, Anna Cusco, Yaozhong Zhang, Juan Salvador Inda-Diaz, Chengkai Zhu, Alexandre Areias Castro, Xinrun Yang, Jiabao Yu, Gaofei Jiang, Xing-Ming Zhao, Luis Pedro Coelho in bioRxiv doi:10.64898/2026.03.20.71308
People
Angelica Jara is a postdoc researcher from the Institut de Biologia Evolutiva (IBE) in Barcelona, who will be visiting the lab in April and May. Her research focuses on studying the function and evolutionary history of secondary chromosomes in non-model bacteria. Find out more: Angelica’s LinkedIn | IBE Evolutionary Microbiology Group.
Papers
Confidence-based prediction of antibiotic resistance at the patent level, by Juan, has been published in mBio. This paper presents a transformer‑based deep learning approach that combines AST data and patient information to predict antimicrobial resistance even in untested antibiotics.
The microbiome-kidney-heart axis paper co-authored by Luis has been published in Nature Communications. The study investigated whether gut microbiome derived metabolites can serve as early predictors of future cardiovascular diseases in healthy Europeans.
Conferences and presentations
A workshop led by Juan is accepted for EDAR8. We will also hold the internal SEARCHER Consortium meeting immediately after EDAR.
Luis presented a talk titled “AI and big data in microbiology: the hype, the promise, and the disappointments” at BrisJAMS.
Tools
Jug is updated to v2.5.0. This update provides faster Polars DataFrame storage, more flexible configuration via project‑local .jugrc files. It also introduces installation support for the jug assistant skill in Codex and Claude code, along with several Python3 bug fixes.
GMSC-mapper v0.2.0 released. GMSC-mapper aligns small proteins to the GMSC. This update brings smaller database downloads (compressed indexes decompressed on the fly), version-stamped outputs for reproducibility, and several bug fixes including a contig length check and duplicate sequence filtering. Note that DIAMOND and MMseqs2 must now be pre-installed.
SPIREpy (which enables access to SPIRE) v0.2.0 has been released. This update introduces changes to support the transition to SPIRE2, along with performance enhancements such as improved storage caching. It also expands functionality with a new suite of tools for downloading and interacting with additional database content, including genes, proteins, and assemblies.
Other
Besides the science, we recently went out for a group dinner!

This was mentioned in the 2023 Summer newsletter with a photo of the sampling.



❤︎